Rview
Usefulness of Rview
Rview is an image registration and analysis tool written by
Dr. Colin Studholme, that enables simple calculation of
ictal-interictal SPECT difference images coregistered with an
MRI scan.
Rview does not smooth, warp, or crop the results (since it
does not perform statistics in comparison with a normal
database). For this reason, it is a very useful way to check
SPM results.
For example, Rview can be used to determine if large amplitude
artifacts from outside the brain are creating spurious signals
to appear in the SPM results, or that other problems have not
occurred during the SPM analysis. Rview (or a similar rapid
analysis method) should be used as a double check on all SPM
analyses, and is available as
freeware.
Rview Terminology
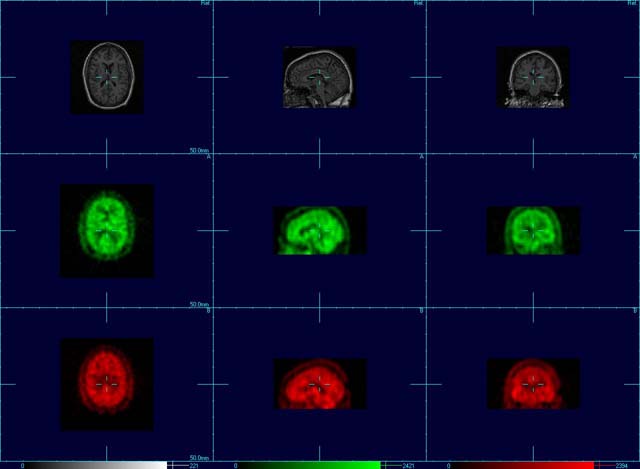
- Ref image (top row of images) = MRI
- Floating Image A (middle row of images) = Interictal SPECT scan
- Floating Image B (bottom row of images) = Ictal SPECT scan
Creating setup files in RView
- Open RView.
- You will see a blank page with 9 panels (three rows of three panels).
- Obtain a high-resolution, coronal, T1-weighted MRI scan, an interictal and an ictal SPECT scanall from the same patient.
(If an MRI is not available, you can still determine differences using just an interictal and ictal SPECT scan using
the interictal SPECT as an anatomical reference, although an MRI is obviously better!)
- Drag and drop the MRI (or interictal SPECT) into the top row of the RView panels.
- Dragging the mouse along the white bar at the bottom left-hand corner of the page can change the contrast of this image.
- Drag and drop the interictal scan into the middle row of the RView page.
- This image is referred to as Floating Image A.
- Dragging the mouse along the red bar in the middle of the bottom of the page can change the contrast of this image.
- Drag and drop the ictal scan into the bottom row of the RView page. This image is referred to as Floating Image B.
- Dragging the mouse along the green bar at the bottom right-hand corner of the page can change the contrast.
- To check the details of the scans, go to Scan Info under the Ref Image (MRI) or Floating Image (SPECT) menus.
-
Align the interictal SPECT scan to the MRI. Go to Floating Images->Image A->Alignment Tool.
Click on the Transaxial Floating Image to Coronal Reference Image button (the upper left-hand button in the Setup Approximate Rotations section), or other buttons as needed to ensure the MRI and interictal SPECT are displayed in the same orientation.
The top two rows of images (MRI and interictal SPECT scans) should now all match in terms of image orientation.
Then, click on Align with Ref Image in the Start Automated Align section.
This alignment step will take about 5 minutes (or less) depending on the speed of your computer.
The program will say busy aligning Image A along the bottom of the screen.
-
When Image A has been aligned to the Reference Image, align the ictal SPECT scan to the interictal SPECT scan.
Go to Floating Images->Image B-> Alignment Tool.
Click on the Transaxial Floating Image to Coronal Image button, or other buttons as needed
to ensure that Floating Image B is in the same orientation as Image A. All three rows of images should now match in terms of image orientation. Then, click on Align with Floating Image A. Again, the program will take a few minutes with this alignment. The bottom of the screen will now read busy aligning Image B.
-
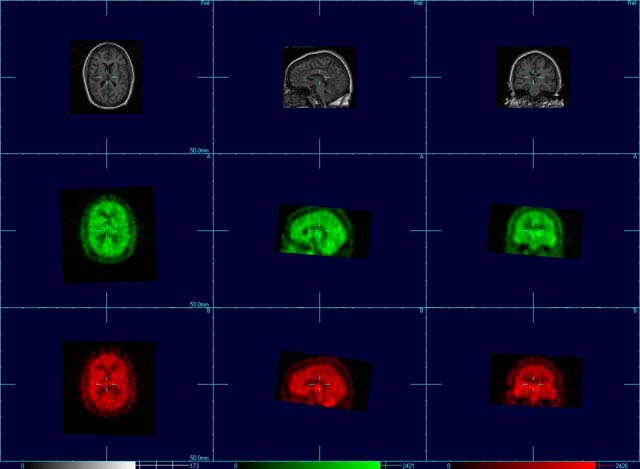
After Image B has been aligned to Image A, the next step is to normalize the intensity of Image B to Image A, to correct for global differences in total brain counts.
Go to Floating Images->Image B->Normalise Image Values to A->Global Normalisation
-
You are ready to view the ictal-interictal differences.
There are a couple of ways you can view the scans.
To see a number of MRI sections at once, go to View Layout->Application Specific Displays->MR-Dual SPECT/PET->ROI Drawing on Multislice Overlay.
To see one set of MRI sections at once that you can scroll through, go to View Layout->Application Specific Displays->MR-Dual-SPECT/PET->ROI Drawing on MRI with Diff Overlay->Transaxial 6.0mm.
To change the color scheme, go to Floating Image->Image B->Colour Tool
(we usually use the sixth choice from the bottom = More Complex II, which allows both increases and decreases in CBF to be visualized on the same images), and Apply to Image.
By left clicking anywhere on the image, you can recenter the display in all three planes to that point.
You can check the registration of the ictal and interictal images with the MRI scan and with each other by going back to View Layout->Application Specific Displays->MR-Dual-SPECT/PET->Ortho-Adj: MR+SPECT A+SPECT B and clicking at various points on the images.
To change the size or location of the images, go to Options->Zoom.
-
To save the setup file, go to View Layout->Save View Setup to..., and you are all set to save your file.
The setup file will allow you to redisplay the images exactly as you had them last time they viewed with Rview.
Multiple setup files can be made for any set of images, viewed in different ways.
Alternatively, Bitmap images can be saved for importation into other software using View Layout->Save as Picture File (bmp).
-
To measure the percent increase or decrease from interictal (image A) to ictal (image B),
draw a region of interest in any plane by right clicking the mouse button and holding it down while drawing.
The percent changes will be displayed below the images.
-
When you want to look at the setup file again, open RView, and drag and drop the setup file
onto the screen.
Checking your results
Compare the results you obtain in RView with those obtained in ISAS to ensure that
the main regions of SPECT increases and decreases are in the same parts
of the brain with both analysis methods.Note that since RView
does not smooth the data, results will appear as more punctuated/scattered
foci, while in ISAS the changes (increases or decreases) will
appear to involve larger brain regions.However, the overall location and
distribution of the most significant increases and decreases should be the same.
In addition, RView should be used to check for artifacts outside the
brain that may introduce spurious signals into the ISAS results. This can
happen, for example, if a large signal increase (or decrease) occurs
just outside the brain, which can be seen with RView (since the images
are not cropped). In ISAS, during the spatial normalization step,
a large signal just outside the brain may be warped into the brain, and
then smoothed into the brain, causing spurious focal changes in that
region. Therefore, if a region shows focal changes in ISAS but no
changes in RView, and if RView shows large adjacent changes outside the
brain, then the ISAS results in that region should be rejected.
In cases where problems arise from large amplitude artifacts outside
the brain, it may be possible to correct the problem by masking out
the artifact before proceeding with the spatial normalization. For
details on how to do this step for SPM2 see:
http://www.psychology.nottingham.ac.uk/staff/cr1/mritut.html#creatingROI
Back to Home
||
Jump to Top
||
Forward to Interpreting Results





